Satpal Virdee and his research group in the MRC PPU have uncovered a striking new role for a family of enzymes in a vital pathway that regulates cell activity and underpins our understanding of health and disease - the result of nearly a decade of collective and sustained scientific efforts to understand the ubiquitin pathway.
Ubiquitin (which takes its name from the adjective “ubiquitous”) is a small protein that can regulate almost any cellular activity in humans, animals and plants. Understanding ubiquitin biology is, therefore, central to our understanding of health and – when something goes wrong – diseases such as cancer and neurodegenerative diseases.
Until recently, it was understood that ubiquitin attaches to a substrate protein (a process called ubiquitylation) through lysine residues to regulate cell activity and that this process can be reversed by the action of deubiquitinating enzymes (DUBs).
When work from the Virdee lab and others indicated that additional amino acids, including serine, threonine and cysteine, are also involved in ubiquitylation [1], Satpal Virdee and his research group in the MRC PPU decided to explore whether any DUBs might target these unusual sites.
“The first stage of our research was very explorative and a number of hypotheses were on the table. We wanted to discover whether ubiquitin could also be removed from serine and threonine residues and, if so, what DUBs were in charge of this.”
Virginia De Cesare, Postdoctoral Researcher with Satpal Virdee
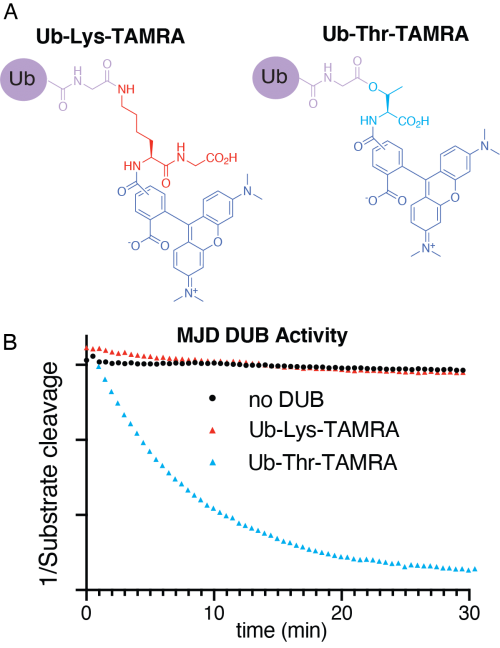
Using chemical methods, the group developed representative model substrates that were used to screen 53 DUBs for non-lysine activity, providing important insights into DUB function.
Strikingly, the researchers identified that a poorly studied family of DUBs - the Machado-Joseph disease family (MJD) – specifically remove ubiquitin from threonine and serine. Ataxin-3, one member of this family of DUBs, is known to be the cause of an inherited neurodegenerative disease (the Machado-Joseph disease, hence the name of the family) whilst another member, JOSD1, is associated with poor gynecological and liver cancer prognosis.
These findings suggest that non-lysine ubiquitination rivals the regulatory sophistication of its conventional counterpart and might serve distinct cellular functions – uncovering new therapeutic targets to treat diseases.
“We now know there are ubiquitin enzymes that specifically favour the attachment to and removal from non-lysine residues. This opens a new dimension to the ubiquitin system that should improve our understanding of both health and disease. The defining characteristics of these enzymes, and the chemical tools we developed, might also be exploitable for the development of new medicines.”
Satpal Virdee, Principal Investigator
This research is the result of nearly a decade of collective and sustained scientific efforts to understand the ubiquitin pathway, with this most recent project taking place from 2017 - 2020. It was published, highlighted and featured on the front cover of PNAS on 26 January 2021.
The research was made possible thanks to the range of reagents, expertise and technologies available within the MRC PPU, MRC PPU Reagents and Services and the University of Dundee’s School of Life Sciences. The group used a unique MALDI-TOF based platform to test the DUB activity – a high-throughput approach that employs nano-liter pipetting system and fast MALDI-TOF MS detection (published in Nature Protocols).
The research was funded by the Medical Research Council, the Biotechnology and Biological Sciences Research Council and The Michael J. Fox Foundation. The research group would also like to acknowledge pharmaceutical companies supporting the Division of Signal Transduction Therapy (Boehringer-Ingelheim, GlaxoSmithKline, and Merck KGaA).
1. McClellan, A.J., S.H. Laugesen, and L. Ellgaard, Cellular functions and molecular mechanisms of non-lysine ubiquitination. Open Biol, 2019. 9(9): p. 190147.
---------------------
News story written by Sarah Patrick, Freelance Communications & Public Engagement Consultant. Published 5 February 2021.

