My lab is interested in understanding a fundamental pathway known as endoplasmic reticulum-specific autophagy, or ER-phagy. The ER is one of the largest organelles that performs a myriad of important functions and is tightly controlled by sophisticated quality control pathways, including the ER-associated degradation (ERAD) and unfolded protein response pathway. In recent years, it has been shown that damaged or superfluous ER can also be specifically degraded via ER-phagy. Clinical mutations of ER-phagy regulators are associated with pathologies such as neurodevelopment defects, pancreatic stress, and cancer metastasis. The field has made significant progress in characterising the ERAD and UPR pathways. However, beyond the handful of ER surface receptors that serve as ‘eat-me’ signals during ERphagy, our understanding of the physiological importance of ERphagy and its disease relevance is still in its infancy.
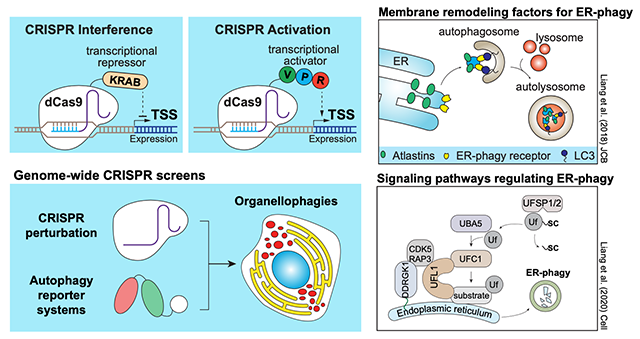
My lab uses state-of-the-art genome-wide CRISPRi and CRISPRa screening technology, fluorescence-activated cell sorting (FACS), mass spectrometry, fluorescence microscopy, and various biochemical assays to uncover novel regulators and signalling pathways that regulate ERphagy activity. We developed an ERphagy reporter system (ER-Autophagy Tandem Reporter; EATR) system that allows us to quantitatively measure ERphagy by flow cytometry. This led us to identify Atlastins, a family of ER transmembrane GTPases, as key factors involved in ER membrane remodelling during ERphagy (Liang et al., JCB, 2018). We further performed a genome-wide CRISPRi screen which led to the identification of ER surface UFMylation as a key factor for ERphagy. Disruption of UFMylation perturbs ERphagy and leads to elevated ER stress and upregulation of UPR (Liang et al., Cell 2020).
Our current research focus:
- Understanding the role of ERphagy during cancer metastasis
- Understanding the crosstalk between ERAD, UPR, and ERphagy
- Genome-wide functional screens to explore novel/understudied ER/organelle biologies
ERphagy has been reported to have varying functions/effects at different stages of cancer development. We are currently focusing on understanding the involvement of ERphagy during cancer metastasis in the hopes that our work can better inform on further cancer treatment. This work is funded by a CRUK-Career Development Fellowship. In addition, we are also interested in understanding the regulation of ER-phagy by UPR. Exactly WHY and HOW cells employ both ERAD (proteasomal pathway) and ERphagy (autolysosomal pathway) to tackle ER stress remains unclear. We intend to perform genome-wide functional screens and ‘-omic’ studies to understand the pathways that regulate the crosstalk between ERAD, UPR, and ERphagy. Last but not least, our lab is also interested in employing genome-wide CRISPRi and CRISPRa screens to explore other under-characterized ER-related biologies. Our lab also has on-going collaborations with other groups with similar interests in other organelle homeostasis and selective autophagy.
Values
The Liang Lab strongly believes in strength and synergy from diversity. We welcome applications from students and postdocs of different ethnicities, nationalities, LGBTQIA+ representation, and disability status. We are committed to building a lab culture that ensures creative thinking, openness in science, and mutual respect. If you are interested in joining the team, please get in touch!



